Drug Discovery
 High throughput screening is the process where a large chemical library is screened for activity in an assay, typically using 384 or 1536-well plate technology. The Penn Center for Molecular Discovery (PCMD) is one of nine centers participating in the NIH Molecular Libraries Screening Centers Network (MLSCN) to screen the NIH repository for biological activity in assays submitted by scientists around the country. High throughput screening is the process where a large chemical library is screened for activity in an assay, typically using 384 or 1536-well plate technology. The Penn Center for Molecular Discovery (PCMD) is one of nine centers participating in the NIH Molecular Libraries Screening Centers Network (MLSCN) to screen the NIH repository for biological activity in assays submitted by scientists around the country.
Miniaturization technology allows scientists to screen thousands of different molecules using tiny wells in plastic plates. But the decreasing size of these wells reached a practical limit of a few microliters. The Diamond laboratory invented a method that doesn’t use wells at all. Molecules are printed like ink as individual nanoliter droplets on a glass microscope slide. An entire library of 100,000 compounds now fits in a slide holder the size of a deck of cards. Using piezo-aerosol delivery methods, Diamond can then add various enzymes, reactants, and fluorescent detectors to each nanoliter reaction. First published in PNAS in 2003, the scale-down saves costs on reagents and robotics needed to move multiple plates around; and, like a printing press, they can print as many copies as they need to use in different screening campaigns.
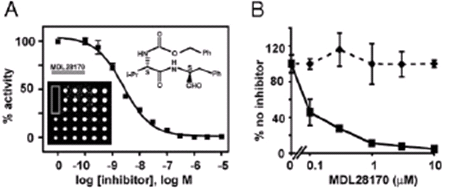
Diamond’s lab has already put the new HTS process to use. This year, Penn microbiologist Paul Bates identified a target, an enzyme called cathepsin L, required for the SARS (severe acute respiratory syndrome) virus to infect cells. Diamond, while independently screening for enzyme inhibitors, found one called MDL28170 that potently inhibited cathepsin L. Bates then tested the compound back in his biological assay with the SARS virus; sure enough, the molecule prevented the infection processes, giving science a new tool for the development of therapeutics against the SARS virus (Simmons et al. PNAS 2005 ).
Publications
| 1. |
2003 |
Gosalia DN, Diamond SL. Printing chemical libraries on microarrays for fluid phase nanoliter reactions. Proc Natl Acad Sci U S A 2003; 100 (15): 8721-8726  |
| 2. |
2005 |
Ma H, Horiuchi KY, Wang Y, Kucharewicz SA, Diamond SL. Nanoliter homogenous ultra-high throughput screening microarray for lead discoveries and IC50 profiling. Assay Drug Dev Technol 2005; 3 (2): 177-187  |
| 3. |
|
Simmons G, Gosalia DN, Rennekamp AJ, Reeves JD, Diamond SL, Bates P. Inhibitors of cathepsin L prevent severe acute respiratory syndrome coronavirus entry. Proc Natl Acad Sci U S A 2005; 102 (33): 11876-11881  |
| 4. |
2006 |
Horiuchi KY, Wang Y, Diamond SL, Ma H. Microarrays for the functional analysis of the chemical-kinase interactome. J Biomol Screen 2006; 11 (1): 48-56  |
| 5. |
2007 |
Myers MC, Napper AD, Motlekar N, Shah PP, Chiu CH, Beavers MP, Diamond SL, Huryn DM, Smith AB. Identification and characterization of 3-substituted pyrazolyl esters as alternate substrates for cathepsin B: The confounding effects of DTT and cysteine in biological assays. Bioorganic & Medicinal Chemistry Letters 2007; 17 (17): 4761-4766  |
| 6. |
2008 |
Beavers MP, Myers MC, Shah PP, Purvis JE, Diamond SL, Cooperman BS, Huryn DM, Smith AB, 3rd. Molecular docking of cathepsin L inhibitors in the binding site of papain. J Chem Inf Model 2008; 48 (7): 1464-1472  |
| 7. |
|
de Almeida RA, Burgess D, Shema R, Motlekar N, Napper AD, Diamond SL, Pavitt GD. A Saccharomyces cerevisiae cell-based quantitative beta-galactosidase assay compatible with robotic handling and high-throughput screening. Yeast 2008; 25 (1): 71-76  |
| 8. |
|
Huang AH, Motlekar NA, Stein A, Diamond SL, Shore EM, Mauck RL. High-throughput screening for modulators of mesenchymal stem cell chondrogenesis. Ann Biomed Eng 2008; 36 (11): 1909-1921  |
| 9. |
|
Motlekar N, Diamond SL, Napper AD. Evaluation of an orthogonal pooling strategy for rapid high-throughput screening of proteases. Assay Drug Dev Technol 2008; 6 (3): 395-405  |
| 10. |
|
Myers MC, Shah PP, Beavers MP, Napper AD, Diamond SL, Smith AB, 3rd, Huryn DM. Design, synthesis, and evaluation of inhibitors of cathepsin L: Exploiting a unique thiocarbazate chemotype. Bioorg Med Chem Lett 2008; 18 (12): 3646-3651  |
| 11. |
|
Shah PP, Myers MC, Beavers MP, Purvis JE, Jing H, Grieser HJ, Sharlow ER, Napper AD, Huryn DM, Cooperman BS, Smith AB, 3rd, Diamond SL. Kinetic characterization and molecular docking of a novel, potent, and selective slow-binding inhibitor of human cathepsin L. Mol Pharmacol 2008; 74 (1): 34-41  |
| 12. |
|
Wong EY, Diamond SL. Enzyme microarrays assembled by acoustic dispensing technology. Anal Biochem 2008; 381 (1): 101-106  |
| 13. |
2009 |
Pagano N, Wong EY, Breiding T, Liu H, Wilbuer A, Bregman H, Shen Q, Diamond SL, Meggers E. From imide to lactam metallo-pyridocarbazoles: distinct scaffolds for the design of selective protein kinase inhibitors. J Org Chem 2009; 74 (23): 8997-9009  |
| 14. |
|
Wong EY, Diamond SL. Advancing microarray assembly with acoustic dispensing technology. Anal Chem 2009; 81 (1): 509-514  |
| 15. |
2010 |
Brey DM, Motlekar NA, Diamond SL, Mauck RL, Garino JP, Burdick JA. High-throughput screening of a small molecule library for promoters and inhibitors of mesenchymal stem cell osteogenic differentiation. Biotechnol Bioeng 2010; 108 (1): 163-174  |
| 16. |
|
Bucki R, Leszczynska K, Byfield FJ, Fein DE, Won E, Cruz K, Namiot A, Kulakowska A, Namiot Z, Savage PB, Diamond SL, Janmey PA. Combined antibacterial and anti-inflammatory activity of a cationic disubstituted dexamethasone-spermine conjugate. Antimicrob Agents Chemother 2010; 54 (6): 2525-2533  |
| 17. |
|
Clifton JD, Lucumi E, Myers MC, Napper A, Hama K, Farber SA, Smith AB, 3rd, Huryn DM, Diamond SL, Pack M. Identification of novel inhibitors of dietary lipid absorption using zebrafish. PLoS One 2010; 5 (8): e12386  |
| 18. |
|
Gosalia D, Diamond SL. Nanodroplet chemical microarrays and label-free assays. Methods Mol Biol 2010; 669 : 69-78  |
| 19. |
|
Liu Z, Myers MC, Shah PP, Beavers MP, Benedetti PA, Diamond SL, Smith AB, 3rd, Huryn DM. Design, synthesis and biological evaluation of a library of thiocarbazates and their activity as cysteine protease inhibitors. Comb Chem High Throughput Screen 2010; 13 (4): 337-351  |
| 20. |
|
Lucumi E, Darling C, Jo H, Napper AD, Chandramohanadas R, Fisher N, Shone AE, Jing H, Ward SA, Biagini GA, DeGrado WF, Diamond SL, Greenbaum DC. Discovery of potent small-molecule inhibitors of multidrug-resistant Plasmodium falciparum using a novel miniaturized high-throughput luciferase-based assay. Antimicrob Agents Chemother 2010; 54 (9): 3597-3604  |
| 21. |
|
Parussini F, Coppens I, Shah PP, Diamond SL, Carruthers VB. Cathepsin L occupies a vacuolar compartment and is a protein maturase within the endo/exocytic system of Toxoplasma gondii. Mol Microbiol 2010; 76 (6): 1340-1357  |
| 22. |
|
Shah PP, Wang T, Kaletsky RL, Myers MC, Purvis JE, Jing H, Huryn DM, Greenbaum DC, Smith AB, 3rd, Bates P, Diamond SL. A small-molecule oxocarbazate inhibitor of human cathepsin L blocks severe acute respiratory syndrome and ebola pseudotype virus infection into human embryonic kidney 293T cells. Mol Pharmacol 2010; 78 (2): 319-324  |
| 23. |
|
Sivendran S, Jones V, Sun D, Wang Y, Grzegorzewicz AE, Scherman MS, Napper AD, McCammon JA, Lee RE, Diamond SL, McNeil M. Identification of triazinoindol-benzimidazolones as nanomolar inhibitors of the Mycobacterium tuberculosis enzyme TDP-6-deoxy-d-xylo-4-hexopyranosid-4-ulose 3,5-epimerase (RmlC). Bioorg Med Chem 2010; 18 (2): 896-908  |
| 24. |
2011 |
Huang F, Motlekar NA, Burgwin CM, Napper AD, Diamond SL, Mazin AV. Identification of specific inhibitors of human RAD51 recombinase using high-throughput screening. ACS Chem Biol 2011; 6 (6): 628-635  |
| 25. |
|
LaFleur MD, Lucumi E, Napper AD, Diamond SL, Lewis K. Novel high-throughput screen against Candida albicans identifies antifungal potentiators and agents effective against biofilms. J Antimicrob Chemother 2011; 66 (4): 820-826  |
| 26. |
|
Moorwood C, Lozynska O, Suri N, Napper AD, Diamond SL, Khurana TS. Drug discovery for Duchenne muscular dystrophy via utrophin promoter activation screening. PLoS One 2011; 6 (10): e26169  |
| 27. |
|
Outten JT, Cheng X, Gadue P, French DL, Diamond SL. A high-throughput multiplexed screening assay for optimizing serum-free differentiation protocols of human embryonic stem cells. Stem Cell Res 2011; 6 (2): 129-142  |
| 28. |
2012 |
Outten JT, Gadue P, French DL, Diamond SL. High-throughput screening assay for embryoid body differentiation of human embryonic stem cells. Curr Protoc Stem Cell Biol 2012; Chapter 1 : Unit 1D 6  |
| 29. |
|
Tan GS, Chiu CH, Garchow BG, Metzler D, Diamond SL, Kiriakidou M. Small molecule inhibition of RISC loading. ACS Chem Biol 2012; 7 (2): 403-410  |
|





